Using AI to identify the aggressiveness of prostate cancer
Update November 12, 2020: a follow-up research study showing how pathologists can benefit from using this technology was published in JAMA Network Open.
Prostate cancer diagnoses are common, with 1 in 9 men developing prostate cancer in their lifetime. A cancer diagnosis relies on specialized doctors, called pathologists, looking at biological tissue samples under the microscope for signs of abnormality in the cells. The difficulty and subjectivity of pathology diagnoses led us to develop an artificial intelligence (AI) system that can identify the aggressiveness of prostate cancer.
Since many prostate tumors are non-aggressive, doctors first obtain small samples (biopsies) to better understand the tumor for the initial cancer diagnosis. If signs of tumor aggressiveness are found, radiation or invasive surgery to remove the whole prostate may be recommended. Because these treatments can have painful side effects, understanding tumor aggressiveness is important to avoid unnecessary treatment.
Grading the biopsies
One of the most crucial factors in this process is to “grade” any cancer in the sample for how abnormal it looks, through a process called Gleason grading. Gleason grading involves first matching each cancerous region to one of three Gleason patterns, followed by assigning an overall “grade group” based on the relative amounts of each Gleason pattern in the whole sample. Gleason grading is a challenging task that relies on subjective visual inspection and estimation, resulting in pathologists disagreeing on the right grade for a tumor as much as 50 percent of the time. To explore whether AI could assist in this grading, we previously developed an algorithm that Gleason grades large samples (i.e. surgically-removed prostates) with high accuracy, a step that confirms the original diagnosis and informs patient prognosis.
Our research
In our recent work, “Development and Validation of a Deep Learning Algorithm for Gleason Grading of Prostate Cancer from Biopsy Specimens”, published in JAMA Oncology, we explored whether an AI system could accurately Gleason grade smaller prostate samples (biopsies). Biopsies are done during the initial part of prostate cancer care to get the initial cancer diagnosis and determine patient treatment, and so are more commonly performed than surgeries. However, biopsies can be more difficult to grade than surgical samples due to the smaller amount of tissue and unintended changes to the sample from tissue extraction and preparation process. The AI system we developed first “grades” each region of biopsy, and then summarizes the region-level classifications into an overall biopsy-level score.
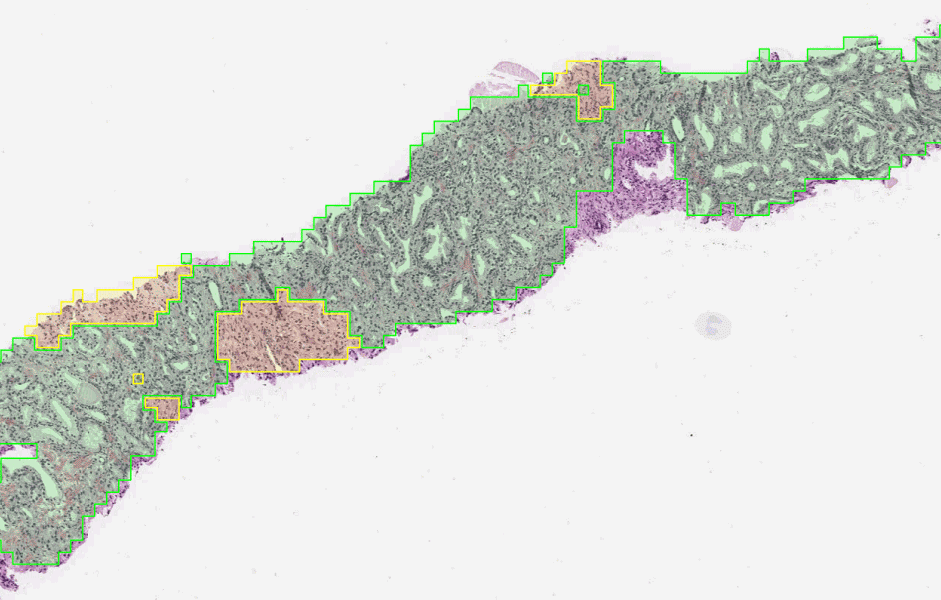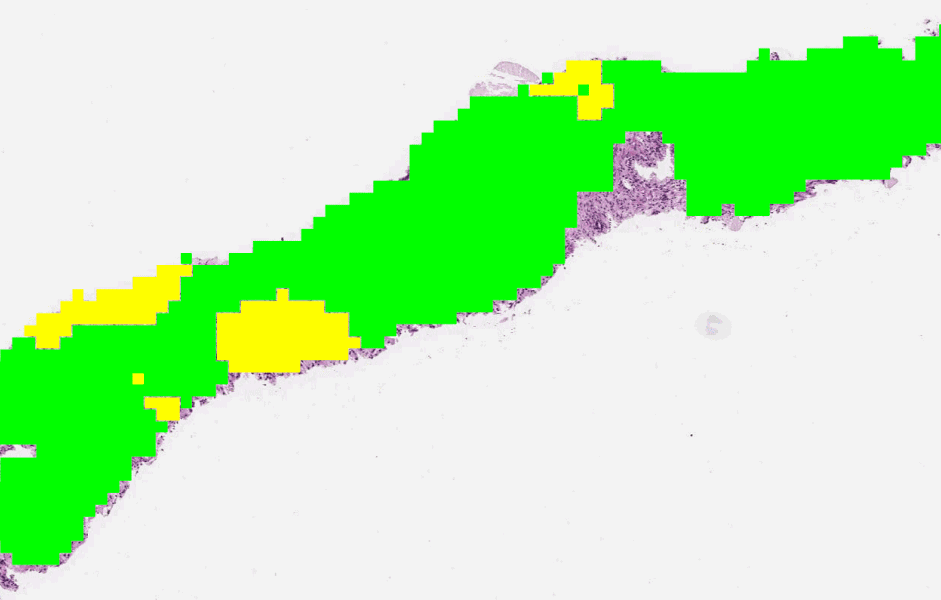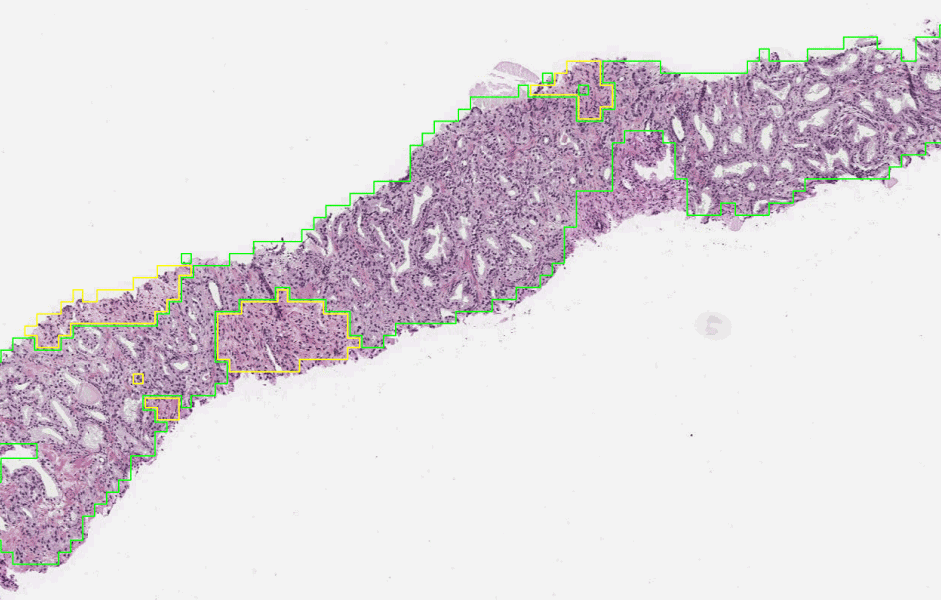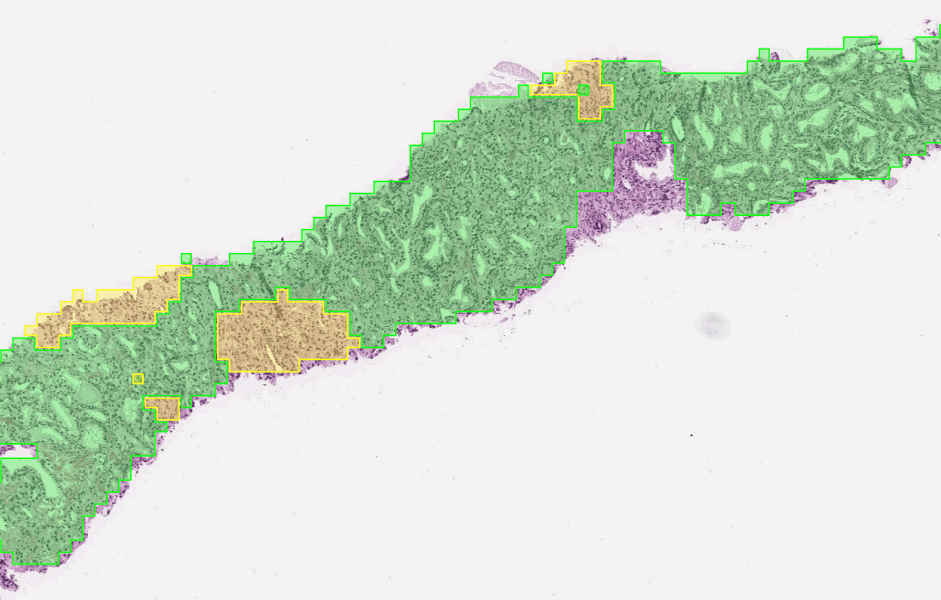
The first stage of the deep learning system Gleason grades every region in a biopsy. In this biopsy, green indicates Gleason pattern 3 while yellow indicates Gleason pattern 4.
Our results
Given the complexity of Gleason grading, we worked with six experienced expert pathologists to evaluate the AI system. These experts, who have specialized training in prostate cancer and an average of 25 years of experience, determined the Gleason grades of 498 tumor samples. Highlighting how difficult Gleason grading is, a cohort of 19 “general” pathologists (without specialist training in prostate cancer) achieved an average accuracy of 58 percent on these samples. By contrast, our AI system’s accuracy was substantially higher at 72 percent. Finally, some prostate cancers have ambiguous appearances, resulting in disagreements even amongst experts. Taking this uncertainty into account, the deep learning system’s agreement rate with experts was comparable to the agreement rate between the experts themselves.
Potential cancer pathology workflow augmented with AI-based assistive tools: a tumor sample is first collected and digitized using a high-magnification scanner. Next, the AI system provides a grade group for each sample.
These promising results indicate that the deep learning system has the potential to support expert-level diagnoses and expand access to high-quality cancer care. To evaluate if it could improve the accuracy and consistency of prostate cancer diagnoses, this technology needs to be validated as an assistive tool in further clinical studies and on larger and more diverse patient groups. However, we believe that AI-based tools could help pathologists in their work, particularly in situations where specialist expertise is limited.
Our research advancements in both prostate and breast cancer were the result of collaborations with the Naval Medical Center San Diego and support from Verily. Our appreciation also goes to several institutions that provided access to de-identified data, and many pathologists who provided advice or reviewed prostate cancer samples. We look forward to future research and investigation into how our technology can be best validated, designed and used to improve patient care and cancer outcomes.